Frontiers in Technology
Poster Session B
(1045-B) A highly automated microscopy analytics pipeline for high-throughput organ-on-chip platforms
Thursday, May 25, 2023
13:30 - 14:30 CET
Location: Hall 3
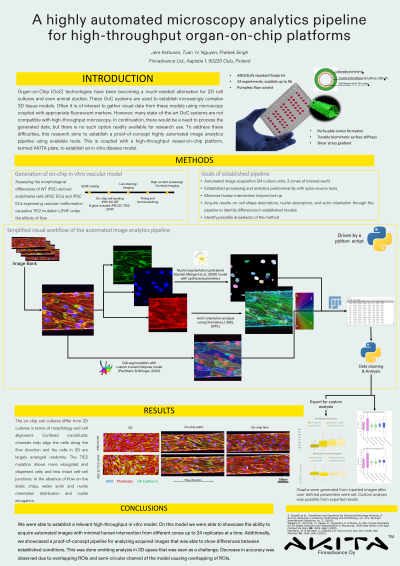

Abstract: In recent years, organ-on-chip (OoC) technology have been becoming a much-needed alternative to conventional 2D cell cultures and even animal studies. OoC systems are a platform to construct complex 3D tissue models that provide relevant in vivo parameters. In these models, the information of interest often pertains to cellular features such as the morphology of the cell or subcellular localization of features of interest. A visual acquisition system, such as confocal microscope, is often used in conjunction with an appropriate fluorescent marker to gather this data. However, many of the current state-of-the-art OoC systems lack compatibility with high-throughput imaging systems. Additionally, many of the available tools for the analysis of acquired visual data seem to lack the necessary capabilities to keep up with more complex tissue models out of the box. To address these difficulties, this research aims to develop a highly automated image analysis pipeline coupled with a high throughput vessel-on-chip platform, termed AKITA Plate, and establish an in vitro disease model with advanced data analytics.
We developed a standardized, high-throughput vessel-on-chip platform capable of 24 parallel experiments on a standardized ANSI 96-well plate footprint that can be integrated into an industrial automation system. This platform was used to establish comparative vascular tissue models between healthy endothelial cells and endothelial cells carrying a vascular anomaly (VA) Tie2 point mutation. These models were subjected to various micro-physiological stimuli before being immunostained with appropriate markers and imaged using a confocal microscope to generate datasets for pipeline assessment. This data was used to evaluate the usability of a variety of available open-source tools and combinations thereafter to produce a proof-of-concept pipeline for the analysis of features of interest such as morphology, orientation, and attachment of cells. This was achieved utilizing available machine learning (ML) models mainly used for the analysis of 2D cell cultures as a starting point to train appropriate models for an OoC use case.
In a comparative study between healthy and mutated cells, we were able to utilize the established pipeline to extract data out of acquired datasets with minimal human intervention. The extracted data of cell shape, orientation, and attachment was able to show an expected difference between healthy and mutation-carrying cells. With this, we have been able to establish a proof-of-concept image analytics pipeline with a minimal human intervention that could be implemented into automated systems with further improvements which can be used to aid in data generation and analysis in high-throughput use cases, such as drug testing.
We developed a standardized, high-throughput vessel-on-chip platform capable of 24 parallel experiments on a standardized ANSI 96-well plate footprint that can be integrated into an industrial automation system. This platform was used to establish comparative vascular tissue models between healthy endothelial cells and endothelial cells carrying a vascular anomaly (VA) Tie2 point mutation. These models were subjected to various micro-physiological stimuli before being immunostained with appropriate markers and imaged using a confocal microscope to generate datasets for pipeline assessment. This data was used to evaluate the usability of a variety of available open-source tools and combinations thereafter to produce a proof-of-concept pipeline for the analysis of features of interest such as morphology, orientation, and attachment of cells. This was achieved utilizing available machine learning (ML) models mainly used for the analysis of 2D cell cultures as a starting point to train appropriate models for an OoC use case.
In a comparative study between healthy and mutated cells, we were able to utilize the established pipeline to extract data out of acquired datasets with minimal human intervention. The extracted data of cell shape, orientation, and attachment was able to show an expected difference between healthy and mutation-carrying cells. With this, we have been able to establish a proof-of-concept image analytics pipeline with a minimal human intervention that could be implemented into automated systems with further improvements which can be used to aid in data generation and analysis in high-throughput use cases, such as drug testing.
- JK
Jere Kettunen
Lead Biofabrication Engineer
Finnadvance
Oulu, Pohjois-Pohjanmaa, Finland