Frontiers in Technology
Poster Session B
(1003-B) Automated and miniaturized 13C-isotopic labeling experiments in microtiter plate scale
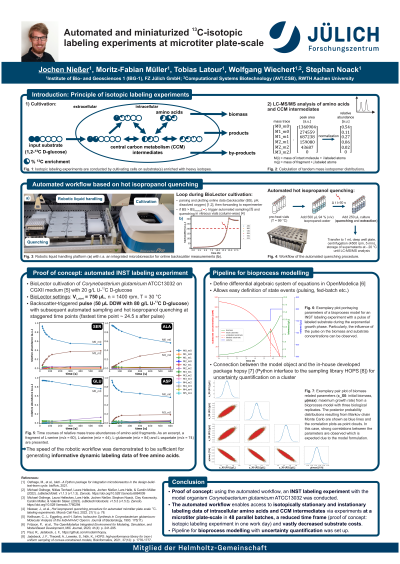

In order to make informed decisions during bioprocess development, the screening of microbial strain variants and the characterization of potential production hosts are of utmost importance. Isotopic labeling experiments (ILE) have numerous applications toward this objective, i.e. 13C metabolic flux analysis (MFA) to elucidate absolute intracellular reaction rates or flux ratio analysis yielding relative pathway usage. Thereby, bottlenecks, avenues for by‑product formation, and other limitations may be recognized and rationally approached. However, the low experimental throughput and high demand of expensive labeled substrate would prohibit such investigations of even the top performing strain variants, let alone a larger selection. Therefore, we have developed and validated an automated experimental workflow for ILEs in microtiter plate‑scale capable of accommodating up to 48 cultivations in parallel.
Development of a quenching method for an automation platform
The present method was designed to enable the investigation of free metabolites at a microliter‑scale and is based on yesteryear’s hot solvent quenching. It utilizes an isopropanol‑water solution heated to its boiling point to quench the metabolism while concomitantly permeating the cell membrane and extracting metabolites in a one‑pot process commonly referred to as whole‑broth sampling. In order to assess whether the inactivation of enzymes leading to the “quenching” effect occurred sufficiently rapid to avoid a systematic bias, the method was tested by spiking the quenching reagent with U13C d-glucose. Since no labeling fraction beyond natural labeling was observed, it was deduced that the quenching method is valid.
Proof of concept for isotopically instationary (INST) labeling experiments
As a proof of concept, an automated INST labeling experiment was conducted. In this case, C. glutamicum ATCC13032 was grown on unlabeled d‑glucose, pulsed with a solution of U13C d‑glucose during the mid‑exponential growth phase, and subsequently sampled in biological triplicates. Currently, the lowest time interval reached between pulsing and quenching amounts to roughly 20 s due to limitations of the robotic platform. Thus, for intermediates of glycolysis and pentose phosphate pathway, merely a stationary endpoint could be recorded since the isotopically instationary phase had already passed. For free amino acids, however, the time course of labeling was duly observed.
Application studies for the automated workflow
Due to the nature of whole‑broth sampling, the measurement of metabolic pool sizes of intermediates is not possible in a straightforward manner. Since pool sizes have traditionally been a prerequisite for 13C‑INST MFA, there have been investigations into model-based in silico pool size estimation.
- JN
Jochen Nießer, M. Sc. (he/him/his)
PhD student
Forschungszentrum Jülich GmbH
Jülich, Nordrhein-Westfalen, Germany