Frontiers in Technology
Poster Session A
(1088-A) New non-toxic dye to probe cell physiology in high content screening
Wednesday, May 24, 2023
13:30 - 14:30 CET
Location: Hall 3
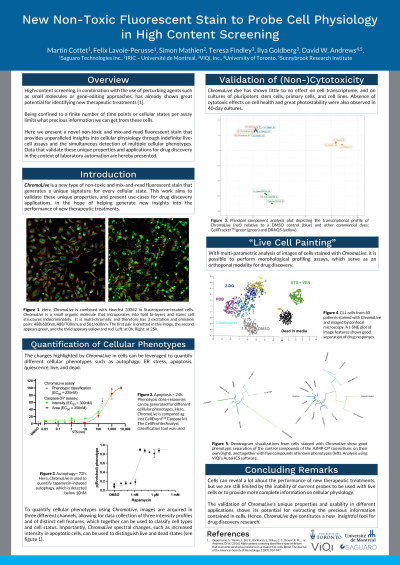
Abstract: High-content imaging approaches, in combination with the use of perturbing agents such as small molecules or CRISPR-driven gene edition, have helped identify new therapeutic compounds [1]. Together with recent advances in image-analysis methods [2], the use of high-content screens is accelerating and therefore making it easier to identify such compounds. However, large-scale high-content screens of live cells have yet to be widely adopted due to the lack of suitable probes compatible with cell growth, as well as limitations of the number of probes that can be used to identify important cellular structures and cellular phenotypes.
Here we present a novel fluorescent non-toxic dye that displays spectral and pattern changes in response to different physiological states and cellular subtypes. With multi-parametric image analysis, its unique morphological and spectral signature allows for the measurement of different cellular phenotypes such as autophagy, ER stress, apoptosis, quiescence and more, with usage of one single dye. Since the dye only gets fluorescent when incorporated into the cell’s membranes, no washing step is needed, therefore making it ideal to include in cell imaging workflows and in large-scale screens. In addition, the dye is non-cytotoxic and does not affect cell proliferation. It has helped provide unparalleled insight into cells cultured in 2D as well as in 3D, and in as diverse as patient derived cancer organoids and stem cell derived hair follicles. More applications are possible with this fluorescent stain, such as morphological profiling screens, and toxicity or genetic screens. The low cytotoxicity and the rich information this dye provides therefore facilitates the assessment of new treatments at more time points and at larger scale, and ultimately paves the way for more discoveries of new therapeutic agents.
[1] Oppermann, S., Ylanko, J., Shi, Y., Hariharan, S., Oakes, C. C., Brauer, P. M., ... & Andrews, D. W. (2016). High-content screening identifies kinase inhibitors that overcome venetoclax resistance in activated CLL cells. Blood, The Journal of the American Society of Hematology, 128(7), 934-947.
[2] Mergenthaler, P., Hariharan, S., Pemberton, J. M., Lourenco, C., Penn, L. Z., & Andrews, D. W. (2021). Rapid 3D phenotypic analysis of neurons and organoids using data-driven cell segmentation-free machine learning. PLoS computational biology, 17(2), e1008630.
Here we present a novel fluorescent non-toxic dye that displays spectral and pattern changes in response to different physiological states and cellular subtypes. With multi-parametric image analysis, its unique morphological and spectral signature allows for the measurement of different cellular phenotypes such as autophagy, ER stress, apoptosis, quiescence and more, with usage of one single dye. Since the dye only gets fluorescent when incorporated into the cell’s membranes, no washing step is needed, therefore making it ideal to include in cell imaging workflows and in large-scale screens. In addition, the dye is non-cytotoxic and does not affect cell proliferation. It has helped provide unparalleled insight into cells cultured in 2D as well as in 3D, and in as diverse as patient derived cancer organoids and stem cell derived hair follicles. More applications are possible with this fluorescent stain, such as morphological profiling screens, and toxicity or genetic screens. The low cytotoxicity and the rich information this dye provides therefore facilitates the assessment of new treatments at more time points and at larger scale, and ultimately paves the way for more discoveries of new therapeutic agents.
[1] Oppermann, S., Ylanko, J., Shi, Y., Hariharan, S., Oakes, C. C., Brauer, P. M., ... & Andrews, D. W. (2016). High-content screening identifies kinase inhibitors that overcome venetoclax resistance in activated CLL cells. Blood, The Journal of the American Society of Hematology, 128(7), 934-947.
[2] Mergenthaler, P., Hariharan, S., Pemberton, J. M., Lourenco, C., Penn, L. Z., & Andrews, D. W. (2021). Rapid 3D phenotypic analysis of neurons and organoids using data-driven cell segmentation-free machine learning. PLoS computational biology, 17(2), e1008630.

Felix Lavoie-Perusse
Co-Founder
Saguaro Technologies
Quebec, QC, Canada