Biology Unveiled
Poster Session A
(1042-A) Towards Analysis of Drug Induced Differential Gene Expression on Droplet Microarray in High Throughput and Nanoliter Format
Wednesday, May 24, 2023
13:30 - 14:30 CET
Location: Hall 3
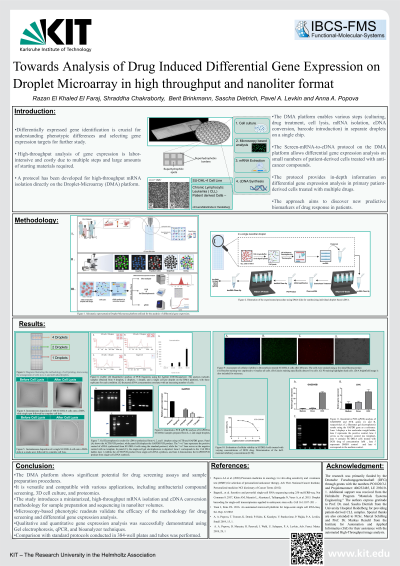

Abstract: The analysis of differential gene expression via mRNA-seq and RT-PCR is a critical tool in identifying the molecular basis for phenotypic differences and selecting gene expression targets for further study. This analysis represents the earliest stimuli-induced changes in cells and provides an accurate molecular footprint of differentially expressed genes under various conditions, enabling the investigation of the mechanism of action of drugs and the identification of biomarkers predictive of drug response. However, performing differential gene expression analysis in a high-throughput manner is both labor-intensive and expensive, as the sample preparation procedure requires multiple single steps, large amounts of starting material, and reagents. To overcome this challenge, we have developed a protocol that enables high-throughput mRNA isolation directly on the Droplet-Microarray (DMA) platform, a highly miniaturized system that enables screening of live cells followed by sample preparation steps including mRNA isolation and conversion to cDNA, to be conducted in individual nanoliter droplets.
The Droplet-Microarray (DMA) platform has the potential to address the aforementioned issue. It is composed of super-hydrophilic spots separated by a superhydrophobic background, resulting in a significant wettability contrast. This platform facilitates the creation of an array of stable nanoliter droplets on super-hydrophilic spots, making it ideal for cell screening purposes. The DMA platform enables the culturing of cells, drug treatment, cell lysis, mRNA isolation, cDNA conversion, and barcode introduction in separate droplets, all on a single DMA chip. As a result, it is an established, high-throughput, and highly miniaturized platform for screening live cells. Multiple droplets can be pooled for amplification, library preparation, and sequencing. Taking advantage of miniaturized volumes, our plan is to apply this protocol to personalized oncology screening, specifically to overcome the challenge of limited availability of primary cells obtained from patient-derived biopsies. Our goal is to establish and evaluate the Screen-mRNA-to-cDNA protocol on the DMA platform in nanoliter volumes and apply it to perform differential gene expression analysis via RT-PCR and mRNA-seq on minute numbers of patient-derived cells treated with anti-cancer compounds on the chip. This will enable us to perform in-depth analysis of drug response in cancer cells from individual patients, conduct research that sheds light on the molecular heterogeneity of drug response among patients, and discover new predictive biomarkers of drug response in patients. DMA is a promising technology with the potential to provide in-depth information about differential gene expression analysis when treating primary patient-derived cells with multiple drugs.
The Droplet-Microarray (DMA) platform has the potential to address the aforementioned issue. It is composed of super-hydrophilic spots separated by a superhydrophobic background, resulting in a significant wettability contrast. This platform facilitates the creation of an array of stable nanoliter droplets on super-hydrophilic spots, making it ideal for cell screening purposes. The DMA platform enables the culturing of cells, drug treatment, cell lysis, mRNA isolation, cDNA conversion, and barcode introduction in separate droplets, all on a single DMA chip. As a result, it is an established, high-throughput, and highly miniaturized platform for screening live cells. Multiple droplets can be pooled for amplification, library preparation, and sequencing. Taking advantage of miniaturized volumes, our plan is to apply this protocol to personalized oncology screening, specifically to overcome the challenge of limited availability of primary cells obtained from patient-derived biopsies. Our goal is to establish and evaluate the Screen-mRNA-to-cDNA protocol on the DMA platform in nanoliter volumes and apply it to perform differential gene expression analysis via RT-PCR and mRNA-seq on minute numbers of patient-derived cells treated with anti-cancer compounds on the chip. This will enable us to perform in-depth analysis of drug response in cancer cells from individual patients, conduct research that sheds light on the molecular heterogeneity of drug response among patients, and discover new predictive biomarkers of drug response in patients. DMA is a promising technology with the potential to provide in-depth information about differential gene expression analysis when treating primary patient-derived cells with multiple drugs.

Razan El Khaled El Faraj
PhD student
Karlsruhe Institute of Technology
Karlsruhe, Baden-Wurttemberg, Germany