Frontiers in Technology
Poster Session B
(1017-B) Long-term proliferation arrayed CRISPR screening by colour competition assay at scale
Thursday, May 25, 2023
13:30 - 14:30 CET
Location: Hall 3
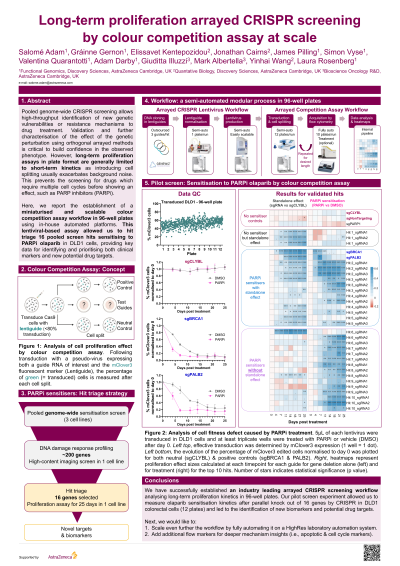
Abstract: Pooled genome-wide CRISPR screening allows high-throughput identification of new genetic vulnerabilities or resistance mechanisms to drug treatment. Validation and further characterisation of the effect of the genetic perturbation using orthogonal arrayed methods is critical to build confidence in the observed phenotype. However, long-term proliferation assays at scale are generally limited to short-term kinetics as introducing cell splitting usually exacerbates background noise and is difficult to automate. This prevents the screening of sensitisation or resistance hits to PARP inhibitors which requires multiple cell cycles before showing an effect.
Here, we report the successful establishment of a miniaturised and scalable growth competition assay workflow in 96-well plates using in-house automated platforms. This lentiviral-based approach measures the cell fitness of fluorescent CRISPR edited cells over non-fluorescent WT cells across multiple rounds of cell splits and data acquisition/analysis by flow cytometry. For example, if loss of a gene has a negative effect on proliferation, a decrease in the edited population of cells will be detected, with varying effect degrees over time depending on the underlying molecular mechanism. The workflow consists of three main independent modules:
(1) high-throughput arrayed production of pseudo-lentiviruses co-expressing a guide RNA of interest and a fluorescent marker,
(2) transduction of cells of interest with those pseudo-viruses at low multiplicity of infection so that only a portion of the cells have been transduced and compete with untransduced cells and,
(3) automated cell split, cell harvesting and PARP inhibitor treatment, and preparation for automated iQue flow cytometer analysis at each time-point (repeated for desired assay time).
We have successfully applied this automated workflow for a pilot screen in colorectal cell line DLD1 stably expressing Cas9. The effect of knocking out 20 pooled screen hits and controls (three guide RNAs per hit) was analysed for 25 days (8 cell splits) with and without PARP inhibition. This pilot provided us with long term kinetic data on both the effect of gene knock-out alone and combined with treatment, which will critically improve our ability to prioritise clinical biomarkers for PARP inhibitors.
Here, we report the successful establishment of a miniaturised and scalable growth competition assay workflow in 96-well plates using in-house automated platforms. This lentiviral-based approach measures the cell fitness of fluorescent CRISPR edited cells over non-fluorescent WT cells across multiple rounds of cell splits and data acquisition/analysis by flow cytometry. For example, if loss of a gene has a negative effect on proliferation, a decrease in the edited population of cells will be detected, with varying effect degrees over time depending on the underlying molecular mechanism. The workflow consists of three main independent modules:
(1) high-throughput arrayed production of pseudo-lentiviruses co-expressing a guide RNA of interest and a fluorescent marker,
(2) transduction of cells of interest with those pseudo-viruses at low multiplicity of infection so that only a portion of the cells have been transduced and compete with untransduced cells and,
(3) automated cell split, cell harvesting and PARP inhibitor treatment, and preparation for automated iQue flow cytometer analysis at each time-point (repeated for desired assay time).
We have successfully applied this automated workflow for a pilot screen in colorectal cell line DLD1 stably expressing Cas9. The effect of knocking out 20 pooled screen hits and controls (three guide RNAs per hit) was analysed for 25 days (8 cell splits) with and without PARP inhibition. This pilot provided us with long term kinetic data on both the effect of gene knock-out alone and combined with treatment, which will critically improve our ability to prioritise clinical biomarkers for PARP inhibitors.
.jpeg.jpg)
Salomé Adam (she/her/hers)
Senior Research Scientist
Astrazeneca Uk Ltd
CAMBRIDGE, England, United Kingdom